Duke University
American Society of Clinical Oncology
American Society of Clinical Oncology
Vrije Universiteit Amsterdam, The Netherlands
Radboud University Medical Center, The Netherlands
Netherlands Cancer Institute, The Netherlands
Queens University, Canada
University of British Columbia, Canada
Queens University, Canada
American Society of Clinical Oncology
nnovations in molecular and genomic analysis have advanced our understanding of cancer biology and progression. It is now clear that cancer is a heterogeneous group of diseases and many of the available therapies may work only in a subset of patients. Furthermore, it is well-established that tailoring treatments to specific oncogenic drivers in patients with various cancers leads to improved clinical outcomes.
Despite remarkable advances in tailoring treatments, the impact of precision medicine has yet to be fully realized in many clinical settings. Nevertheless, this field has been ignited, and we have seen a surge in prospective trials that match patients to drugs based on their tumor’s genomic alterations to identify signals of drug activity.
There have been several successful examples of targeted therapies, including:
- Trastuzumab in breast and gastric cancer patients whose tumors overexpress HER2/neu,
- imatinib and related drugs to target BCR-ABL fusions in chronic myeloid leukemia,
- vemurafenib to treat melanoma patients with tumors that harbor the BRAFV600E mutation, and
- olaparib in patients with ovarian, breast, and pancreatic cancer who have germline alterations in BRCA1/2 as well as in metastatic castration-resistant prostate cancer patients who have a homologous recombination repair deficiency profile.
However, despite these successes and the strong rationale for this approach, there is no assurance that matching drugs to genomic alterations will result in meaningful treatment benefit to patients across all tumor types that harbor the same mutation. This is clearly demonstrated in colorectal cancer patients with the BRAFV600E mutation, where BRAF-directed monotherapy has limited activity compared to its high efficacy in melanoma.
In order to assess the efficacy of targeted therapies outside of their approved indications, several studies were launched in recent years:
- The United States based American Society of Clinical Oncology (ASCO) Targeted Agent and Profiling Utilization Registry (TAPUR) Study (NCT02693535),
- the Drug Rediscovery Protocol (DRUP) Study in the Netherlands (NCT02925234), and
- the Canadian Profiling and Targeted Agent Utilization Trial (CAPTUR) in Canada (NCT03297606).
These trials are designed as phase 2 platform trials where patients whose tumors harbor specific genomic alterations are matched to commercially available therapies known to target those alterations. In each of these trials, cohorts are defined based on the unique combination of targeted therapy (D) administered, genomic alteration (G) in the tumor, and tumor histology (H), and are referred to by D/G/H. Thus, hundreds of cohorts can potentially be formed based on various combinations of D/G/H. These three trials, all employing similar designs and endpoints, have, at present, ten therapies in common that include axitinib, crizotinib, erlotinib, nivolumab plus ipilimumab, olaparib, palbociclib, sunitinib, trastuzumab plus pertuzumab, vemurafenib plus cobimetinib, and vismodegib.
While these studies have collectively enrolled thousands of patients to hundreds of cohorts over the last five years, a major challenge in reporting timely results from these trials is completing cohorts with rare genomic alterations, many of which occur at a frequency of 5 percent or less in the population of patients with cancer. Therefore, the principal investigators of each these trials made a commitment to advancing research through the sharing of trial data. Thus, TAPUR, DRUP, and CAPTUR formed a global collaboration (referred to as TADRUCA), with the main goal of combining data from slowly enrolling cohorts of patients either in the conduct or the analysis stage.
Each of these trials is designed using Simon’s two-stage design. In TAPUR, the null hypothesis is that the probability of objective response or stable disease of at least 16 weeks duration is 15 percent, versus the alternative hypothesis that it is at least 35 percent. Power and Type I error rate are assumed to be 85 percent and one-sided 0.10, respectively. In DRUP and CAPTUR, the null hypothesis is that the probability of objective response is 10 percent, versus the alternative hypothesis that it is at least 30 percent. Power and one-sided Type I error rate are assumed to be 85 percent and 0.078.
Although there are small differences across the three trials in terms of the hypotheses and endpoints, the TADRUCA protocol harmonized the primary endpoint in order to address the primary objective of estimating the objective response rate (ORR) in a cohort (G/D/H) with rare genomic alterations (Table 1).
The TADRUCA protocol utilizes a Simon two-stage approach that tests the null hypothesis of ORR of 10 percent vs. 30 percent (power = 0.85; one-sided type I error rate= 0.10). In the first stage, 8 patients are needed to evaluate the effectiveness of a drug within a cohort. If ≥ 1 patient experiences objective response, 16 more patients are to be included. If ≥5 of 24 patients have objective response, treatment would be considered worthy of further study.
By integrating data across the three trials in either stage 1 or stage 2, cohorts can be completed more rapidly. Data sharing can facilitate the process of evaluating the therapies within a cohort in stage 1. The concept of data sharing during the first stage is a novel paradigm as data across the trials will be used to perform a futility analysis.
Suppose that TAPUR has enrolled five patients in stage 1 in cohort A (e.g., ATM mutation, pancreatic cancer, treated with olaparib) and the cohort enrolled its first patient 20 months ago. By borrowing information from a similar cohort in another study yielding a total of at least eight patients combined, a decision by the Data Safety Monitoring Board (DSMB) can be made to either close or expand that specific cohort.
Alternative Hypothesis
Power
Type I error rate (one-sided)
35%
0.85
0.10
30%
0.85
0.078
30%
0.85
0.078
30%
0.85
0.078
≥2 responses
≥1 response
≥1 response
≥1 response
≥7 responses
≥ 5 responses
≥5 responses
≥5 responses
Lugano (NHL)
IMWG (MM)
iRECIST 1.1(solid & immuno)
RECIST & CA125 (gyno)
RECIST&PSA (prostate)
Lugano (NHL)
IMWG (MM)
*Confirmation of PR and CR is required (for the different types of RECIST criteria) a minimum of 4 weeks after initial documentation.
RECIST&CA125 (gyno)
RANO (glioblastoma)
Lugano (NHL)
IMWG (MM)
**Confirmation of PR and CR is required (≥30 days after the first documentation of PR or CR).
Abbreviations: CA125 = cancer antigen-25 (rising level associated with progression of ovarian cancer); CR = Compete response to treatment per established evaluation criteria; IMWG Criteria = International Myeloma Working Group criteria for the diagnosis of MM; Lugano Criteria = Criteria for classification of treatment response in NHL using 18F-fluorodeoxylucose positron emission tomography; MM = Multiple myeloma; NHL = Non-Hodgkin lymphoma; PD = Progressive disease (no response to treatment) per established evaluation criteria; PR = Partial response to treatment per established evalution criteria; PSA = Prostate specific antigen (rising level and rate of rise associated with prostate cancer progression; RANO Criteria = Response assessment in neuro-oncology criteria (applied to evaluaation of brain tumors such as glioblastoma); RECIST = Response Evaluation Criteria in Solid Tumors; these criteria are based on tumor measurements and include specifications for CR, PR, PD, and SD; iRECIST = RECIST criteria modified for use in evaluating response to immunotherapy; SD = Stable disease
Alternative Hypothesis
Power
Type I error rate (one-sided)
35%
0.85
0.10
30%
0.85
0.078
30%
0.85
0.078
30%
0.85
0.078
or SD at 16 weeks or later.
≥2 responses
≥1 response
≥1 response
≥1 response
≥7 responses
≥ 5 responses
≥5 responses
≥5 responses
Lugano (NHL)
IMWG (MM)
iRECIST 1.1(solid & immuno)
RECIST & CA125 (gyno)
RECIST&PSA (prostate)
Lugano (NHL)
IMWG (MM)
*Confirmation of PR and CR is required (for the different types of RECIST criteria) a minimum of 4 weeks after initial documentation.
RECIST&CA125 (gyno)
RANO (glioblastoma)
Lugano (NHL)
IMWG (MM)
**Confirmation of PR and CR is required (≥30 days after the first documentation of PR or CR).
Abbreviations: CA125 = cancer antigen-25 (rising level associated with progression of ovarian cancer); CR = Compete response to treatment per established evaluation criteria; IMWG Criteria = International Myeloma Working Group criteria for the diagnosis of MM; Lugano Criteria = Criteria for classification of treatment response in NHL using 18F-fluorodeoxylucose positron emission tomography; MM = Multiple myeloma; NHL = Non-Hodgkin lymphoma; PD = Progressive disease (no response to treatment) per established evaluation criteria; PR = Partial response to treatment per established evalution criteria; PSA = Prostate specific antigen (rising level and rate of rise associated with prostate cancer progression; RANO Criteria = Response assessment in neuro-oncology criteria (applied to evaluaation of brain tumors such as glioblastoma); RECIST = Response Evaluation Criteria in Solid Tumors; these criteria are based on tumor measurements and include specifications for CR, PR, PD, and SD; iRECIST = RECIST criteria modified for use in evaluating response to immunotherapy; SD = Stable disease
Data Sharing Implementation
Representatives of each study team formed a Steering Committee that has the overall responsibility to:
- provide advice and make decisions regarding the data sharing protocol, including approving any protocol amendments;
- review and approve abstracts and publications for the different cohorts;
- review and approve data sharing requests (internal or external to the three study teams); and
- provide recommendations about communications regarding the results for combined cohorts.
Although each trial is monitored by an independent DSMB, a streamlined process is planned to approve requests for data sharing expeditiously. Once the Steering Committee approves a proposal for combining a cohort across the three studies, the principal investigator leading the analysis, either in stage 1 or stage 2, will inform their DSMB of the Steering Committee’s approval for combining data across the three studies. A report will be submitted to the DSMB of the group that is leading the analysis to determine whether to close or expand the cohort or release the results.
The goal is to combine data from common cohorts that individual study teams identify as a potential cohort for data sharing. The extent of data shared will depend on the cohort stage: For example, in stage 1 only response data and the total number of patients enrolled are needed, because the only decision to be made is whether the cohort needs to be expanded or closed. On the other hand, for stage 2, other baseline characteristics, prognostic factors, and genomic laboratory information may be needed as this will constitute the final analysis of the cohort. Each study team (CAPTUR, DRUP, and TAPUR) reviews their cohorts on a tri-annual basis and identifies cohorts that are eligible for data sharing. A cohort in a trial will be considered eligible for data sharing (i.e., for combining with a cohort from another trial) if fewer than three patients have been enrolled in the past 12 months, and if the cohort has been opened for at least 24 months (Figure 1). Once the number of patients has been reached for stage 1 (eight patients combined across the studies) or stage 2 (24 patients), data will be combined in a pooled cohort for analysis.
Although pooling data within histology-specific cohorts across the three trials is helpful, for rare variants this strategy may not achieve the target number of patients for either stage 1 or stage 2 analysis. Therefore, collapsing across histology will also be considered at the time when eligible cohorts are evaluated for pooling within cohorts across the three trials. If pooling data within a cohort does not lead to a sample size of eight in stage 1 or 24 patients in stage 2, then cohorts may be considered for grouping across histology until there are at least eight patients in stage 1 (based on the design of the G/D/H data sharing) or 24 patients in stage 2. This sharing strategy among the three studies maximizes the potential to learn quickly without impacting each individual trial’s objectives or goals.
In summary, despite the promise of precision medicine, its clinical application is still limited, and benefits to patients continue to lag. The low number of patients with rare mutations necessitates global collaboration. The TADRUCA collaboration offers a great opportunity to answer questions regarding the potential effectiveness of targeted therapies for cohorts of cancer patients with rare tumor genomic alterations. This data sharing model used during trial enrollment can also be applied to other diseases for faster translation of results and benefit to all patients in the era of precision medicine.
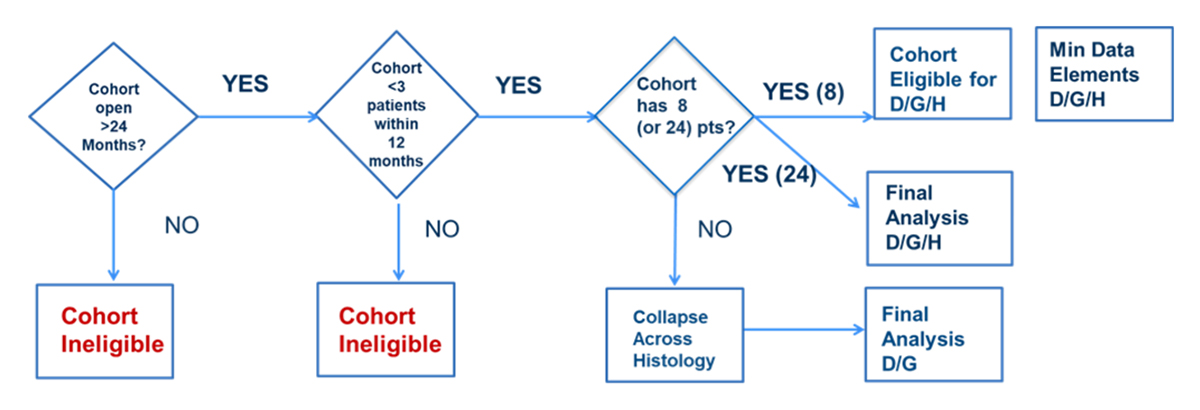
G=Genomic Alteration
Histology=H

